
As spatial biology continues to reshape life science research, selecting the right spatial transcriptomics platform has become a key decision. Whether you’re analyzing FFPE samples, prioritizing subcellular resolution, or exploring high-throughput options, today's commercial solutions—from 10x Genomics to BGI —offer diverse technologies tailored to different experimental needs. In this article, we break down the strengths and technical highlights of leading platforms to help you make the best choice for your research.
Robust and Widely Used for Fresh-Frozen Tissue
Overview:
Visium 1.0, launched in 2019, integrates barcoded capture probes into slide-based spatial arrays. Each slide contains four 6.5 × 6.5 mm capture areas, each with 5,000 barcoded spots (55 μm diameter, 100 μm spacing), making it ideal for broad tissue compatibility and morphological validation.
Strengths:
Supports polyA+ RNA from animals and plants
Compatible with H&E staining on the same section
Works with fresh-frozen and -80°C preserved samples
Proven across multiple species: human, mouse, zebrafish, Arabidopsis, etc.
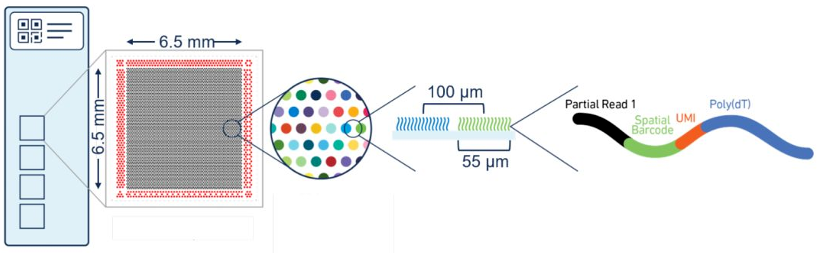
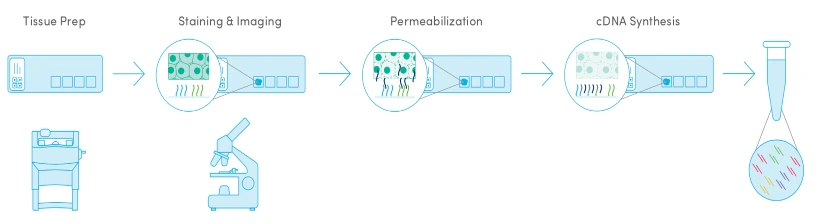
Best for:
General-purpose spatial transcriptomics with moderate resolution and flexible tissue input.
Optimized for FFPE Tissues with Larger Capture Area
Overview:
Visium 2.0 (CytAssist FFPE) builds on Visium 1.0 with redesigned probes that allow FFPE compatibility without requiring polyA tails. A new 11 × 11 mm chip increases spot count to ~14,000 while maintaining the same resolution.
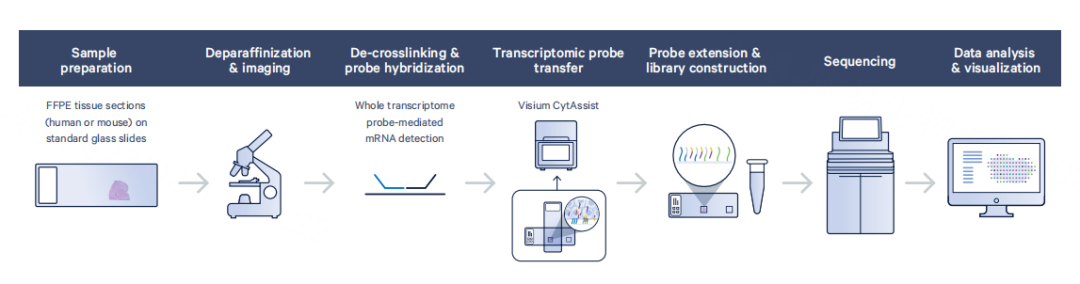
Strengths:
Enables spatial profiling of archival FFPE samples
No permeabilization required—streamlined workflow
H&E stain compatibility retained
Compatible with FFPE, fresh, and frozen samples
Available for human and mouse tissues
Best for:
Researchers working with rare, clinical, or archived FFPE specimens.
High-Resolution Platform with Submicron Spot Spacing
Overview:
Stereo-seq offers spatial transcriptomics with exceptional resolution using dense spot arrays (220 nm diameter, 500 nm spacing). Launched commercially in 2022, it supports both 10×10 mm and 5×5 mm capture chips using poly-dT RNA capture.
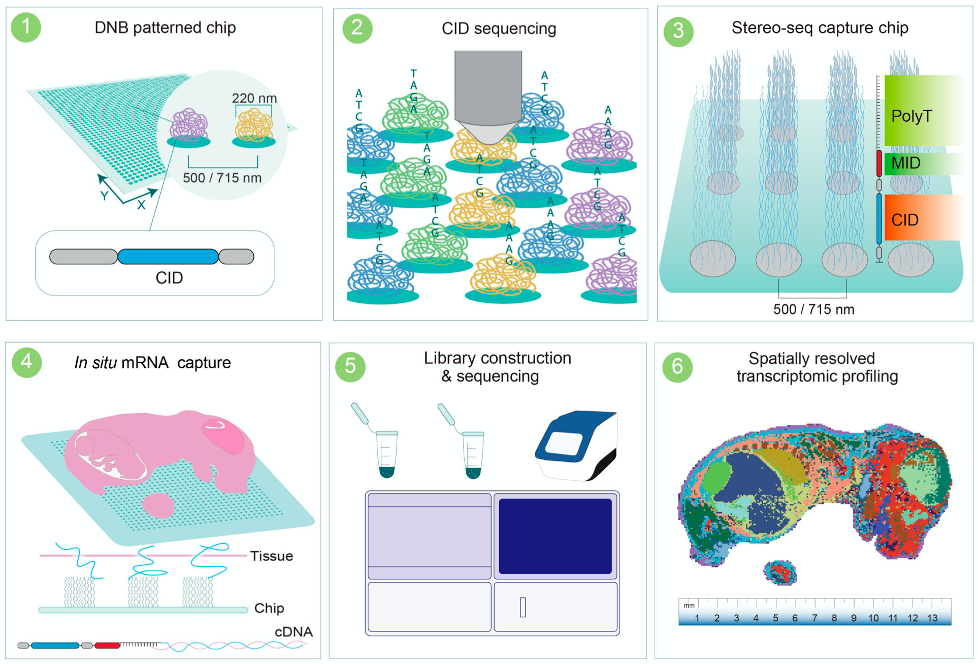
Strengths:
Ultra-high spatial resolution—approaching single-cell level
Broad species and sample compatibility (fresh and -80°C)
Adjustable resolution for tailored analysis
Proven in human, mouse, zebrafish, Arabidopsis, and more
Best for:
Researchers prioritizing fine spatial mapping, single-cell-level resolution, and scalable output.
Targeted Spatial Profiling with Single-Cell Precision
Overview:
Xenium offers high-resolution in situ gene detection via padlock probe technology. Its chip size is 12 × 24 mm, with panels supporting detection of ~300 genes (customizable up to 500). Unlike sequencing-based tools, Xenium provides direct spatial imaging at subcellular resolution.
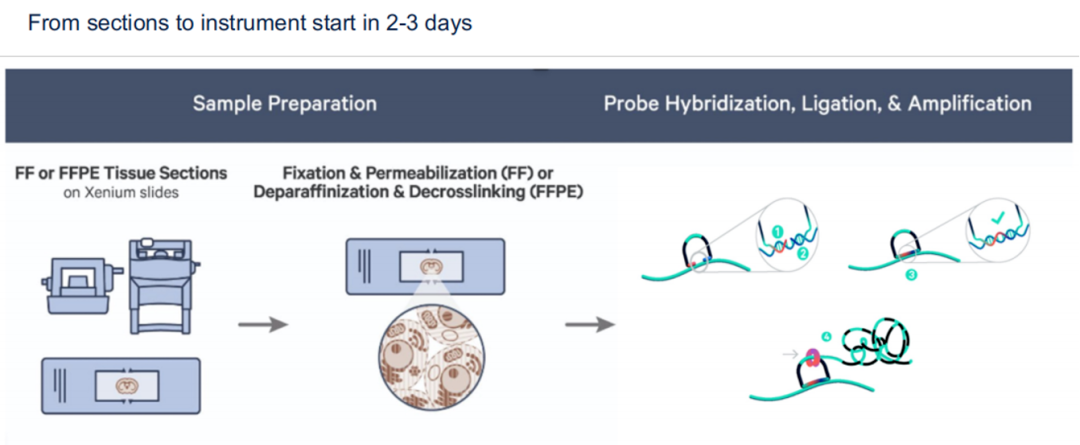
Strengths:
Subcellular-level spatial resolution
Compatible with FFPE, fresh, and frozen tissues
Non-destructive: post-assay H&E and IHC still possible
Targeted panels for cancer and developmental research
Designed for human and mouse tissues
Best for:
Users seeking targeted gene panels with ultra-high-resolution imaging of spatial gene expression.
There is no one-size-fits-all solution in spatial transcriptomics. Each platform offers unique strengths depending on your research priorities:
Research Priority | Best Platform |
FFPE sample compatibility | Visium 2.0 (CytAssist FFPE), Xenium |
Highest spatial resolution | Stereo-seq, Xenium |
Broad gene detection (whole-transcriptome) | Visium 1.0/2.0, Stereo-seq |
Targeted, single-cell-level detection | Xenium |
Broad species/sample coverage | Visium 1.0, Stereo-seq |
With spatial omics still in its growth phase, now is the time to explore new technologies and match the right platform to your experimental goals.
At Omics Empower, we support spatial transcriptomics projects across leading platforms, including 10x Genomics Visium, Xenium In Situ, and STOmics Stereo-seq, helping researchers identify the most suitable workflow for whole-transcriptome discovery, targeted in situ analysis, or ultra-high-resolution spatial profiling.
With more than 500 publications in single-cell sequencing and spatial transcriptomics, our experience spans diverse study goals, sample types, and research applications. Supported by laboratory operations in the U.S., Europe, and Asia, we provide coordinated workflows and reliable delivery for global research teams.
If you are deciding between technologies for your next study, contact us today for a complimentary consultation on platform selection, project feasibility, and workflow design.
If you are exploring spatial transcriptomics platforms for your next project, these related articles may help you better understand Xenium, Visium HD, and Stereo-seq, and compare different workflows based on your study goals and sample requirements:
Beyond Poly(A): Cell Segmentation Joins the 10x Genomics Visium HD Pipeline
Stereo-seq S0.5 Explained: A High-Resolution Option for Small Spatial Samples
Germany: Arnold-Graffi-Haus / D85 Robert-Rössle-Straße 10 13125 Berlin
United States: (CA) 2 Goddard, Irvine, CA 92618
United States: (IL) 8255 Lemont Rd, #1, Darien, IL 60561
Hong Kong: Room 618, Building 6, Hong Kong Science Park, Pak Shek Kok, Hong Kong
Germany: Arnold-Graffi-Haus / D85 Robert-Rössle-Straße 10 13125 Berlin
United States: (CA) 2 Goddard, Irvine, CA 92618
United States: (IL) 8255 Lemont Rd, #1, Darien, IL 60561
Hong Kong: Room 618, Building 6, Hong Kong Science Park, Pak Shek Kok, Hong Kong